
Multilevel omics integration analysis normalized, compared and correlated the batch data from different biomolecular levels, such as genome, transcriptome, proteome, metabolome and lipid, to establish the data relationship between molecules at different levels. Combined with GO function analysis, metabolic pathway enrichment, molecular interactions and other biological functions, the function and regulation mechanism of biomolecules are systematically and comprehensively analyzed.
Multilevel omics integration analysis normalized, compared and correlated the batch data from different biomolecular levels, such as genome, transcriptome, proteome, metabolome and lipid, to establish the data relationship between molecules at different levels. Combined with GO function analysis, metabolic pathway enrichment, molecular interactions and other biological functions, the function and regulation mechanism of biomolecules were systematically and comprehensively analyzed.
Creative Proteomics provides integrated metabolomic and other histological analysis services to assist in subsequent in-depth experimental analysis and applications by screening key metabolic pathways as well as proteins, genes, and lipid products.
The study of microorganisms by 16S and metagenomic techniques, combined with metabolomic analysis, allows for a systematic study of microbial roles and functions.
LEARN MOREAnalysis of differential protein bioinformation, analysis of differential metabolite bioinformation, analysis of metabolite-related enzymes, and metabolic pathway-based protein metabolism integration analysis.
LEARN MORE4D-DIA proteome combined with GC-MS and LC-MS technical services to screen key metabolic pathway-related proteins or metabolites and perform integration analysis.
LEARN MOREA large number of differentially expressed genes are obtained from transcriptome and correlated with differential metabolites obtained from metabolome screening. Analyze metabolite-associated enzymes and differential genes, target key pathways with metabolite changes, and construct core regulatory networks.
LEARN MOREGenotype data of population materials are obtained by second-generation sequencing technology, and genome-wide association analysis of metabolites is carried out by combining metabolome data to locate candidate genes related to the control of metabolic phenotypes and to explore the mechanisms of related metabolic pathways.
LEARN MORELncRNA sequencing is used to obtain a large number of differentially expressed LncRNAs for association analysis with the differential metabolites obtained from metabolome analysis. We analyze the regulatory genes as well as the expressed gene changes, target the key pathways with metabolite changes, and construct the core regulatory network.
LEARN MORE| Service | Technology | Instrument/Equipment | Advantages |
|---|---|---|---|
| Genomics Services | DNA Sequencing Techniques | - Illumina HiSeq 4000 (High-throughput sequencer) | Enables rapid generation of large-scale genomic data |
| - Pacific Biosciences Sequel II (Single-molecule sequencer) | Suitable for long-read sequencing | ||
| Proteomics Services | Mass Spectrometry | - Thermo Fisher Orbitrap Fusion (LC-MS, High-resolution mass spectrometer) | Offers high-resolution for precise protein identification and quantification |
| - Waters Synapt G2-Si (LC-MS, High-sensitivity mass spectrometer) | Provides high sensitivity for detecting protein isoforms | ||
| - Bruker timsTOF Pro (High-resolution mass spectrometer) | Delivers ultra-high resolution for detailed protein structural analysis | ||
| Metabolomics Services | Mass Spectrometry | - Agilent 7890B GC-MS (Gas chromatography-MS) | Analysis of volatile metabolites |
| Thermo Fisher Q Exactive Plus (LC-MS, High-sensitivity mass spectrometer) | High sensitivity liquid chromatography-MS | ||
| Waters Xevo G2-XS QTof (LC-MS, High-resolution mass spectrometer) | High mass resolution for metabolite identification and quantification | ||
| Transcriptomics Services | RNA Sequencing Techniques | - Illumina NovaSeq 6000 (High-throughput sequencer) | High-throughput RNA analysis and expression profiling |
| - Nanopore GridION (Single-molecule RNA sequencer) | Detection of RNA modifications, single-molecule RNA sequencing | ||
| - PacBio Sequel IIe (Long-read sequencer) | Full-length transcript identification, long-read sequencing |
Metabolic Network Analysis: Multi-omics approaches allow for the construction of comprehensive metabolic networks. These networks can reveal metabolic interactions, fluxes, and potential regulatory points.
Biomarker Discovery: By combining different omics data, researchers can identify robust biomarkers for diseases or physiological conditions. This has implications for early diagnosis and personalized medicine.
Functional Annotation of Genes and Proteins: Integrative analysis helps in functionally annotating genes and proteins by considering their roles in metabolic pathways and networks.
Understanding Disease Mechanisms: Multi-omics studies provide a holistic view of disease mechanisms, uncovering intricate relationships between genetic variations, gene expression, protein function, and metabolic dysregulation.
Drug Discovery and Development: Integrated omics data can aid in drug target identification, drug repurposing, and assessing the effects of drugs on multiple biological layers.
Nutrition and Health: Multi-omics approaches can elucidate the impact of nutrition on health by analyzing the interplay between dietary components, gut microbiota, host metabolism, and health outcomes.
Environmental Impact: Studying multi-omics responses to environmental factors provides insights into how environmental exposures affect an individual's health and metabolism.
If you want to know more, please contact us. Looking forward to cooperating with you.
Reference
The study focuses on the metabolic changes that occur during the transition from G0 (resting state) to the G1 phase of the cell cycle, with a particular emphasis on how these changes relate to cancer metabolism. Cancer cells often shift their metabolic preferences, favoring glycolysis over mitochondrial metabolism (TCA cycle) for ATP generation. Understanding these metabolic alterations during the cell cycle entry provides valuable insights into cancer cell behavior.
The study uses FL5.12 cells as the experimental model. These cells are undergoing the transition from the resting state (G0) to the proliferative G1 phase. The analysis is performed at multiple time points to capture the dynamic changes in metabolic pathways during this transition.
Pathway Analysis: The research employs a comprehensive approach to analyze metabolic changes. It focuses on eight key metabolic pathways, including the TCA cycle, glycolysis, nucleotide biosynthesis, lipid synthesis, and the urea cycle. This analysis involves studying the expression of enzymes and the levels of metabolites within these pathways.
Proteomics and Metabolomics: To understand the changes in enzyme expression and metabolite levels, the study uses proteomics and metabolomics techniques. Proteomics involves quantifying the levels of various proteins, including enzymes, while metabolomics measures the levels of metabolites, such as small molecules and intermediates in metabolic pathways.
Extracellular Metabolite Profiling: In addition to intracellular changes, the study explores extracellular metabolite profiles in the growth medium. This approach helps identify metabolites that are consumed or released by cells during the cell cycle transition.
Amino Acid Consumption Analysis: A detailed analysis of amino acid consumption is conducted. This includes quantifying the uptake of specific amino acids and normalizing it based on their relative frequencies in proteins. This analysis provides insights into the amino acids most critical for supporting cell growth and protein synthesis during the transition.
The study yields several key findings:
Metabolic Shift: FL5.12 cells exhibit a metabolic shift during the G0 to G1 transition, similar to the metabolic preferences seen in cancer cells. This involves downregulation of the TCA cycle and upregulation of glycolysis, favoring ATP generation through glycolysis.
Nucleotide Biosynthesis: Enzymes involved in de novo nucleotide biosynthesis are upregulated, indicating increased demand for nucleotides to support cell growth and proliferation.
Lipid Biosynthesis: Key enzymes in fatty acid synthesis are upregulated during the G1 phase, suggesting enhanced lipid biosynthesis to support biomass generation.
Amino Acid Consumption: Methionine (Met) consumption is notably high during the G1-S phase transition, aligning with increased protein synthesis and nucleotide biosynthesis. This correlates with the observed patterns in histone methylation, indicating Met's role in epigenetic modifications during the cell cycle.
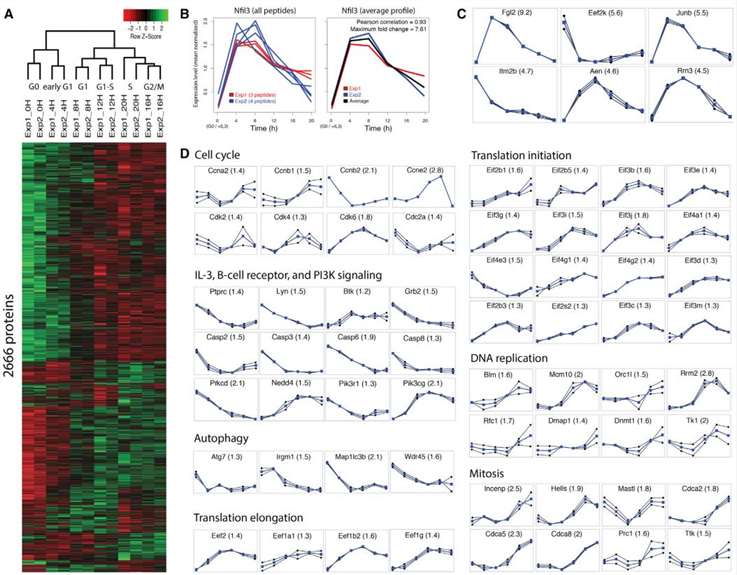 Proteomics Data of Temporal Expression and Highly Confident Individual Proteins
Proteomics Data of Temporal Expression and Highly Confident Individual Proteins
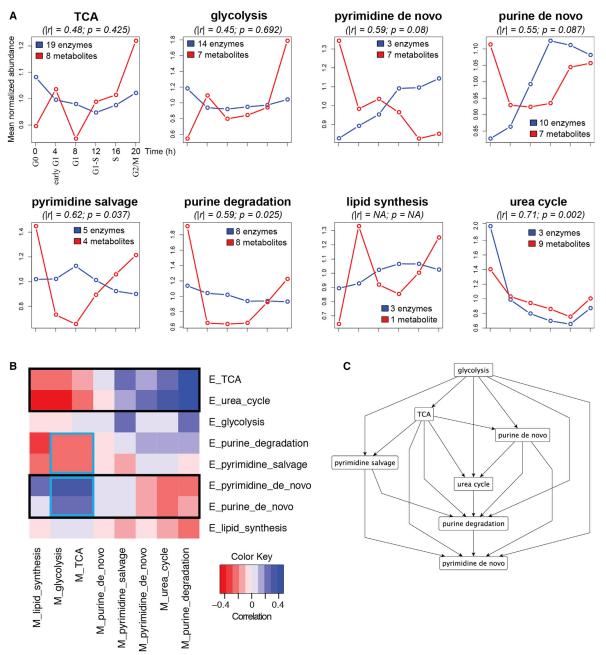 Correlation between Enzymes and Metabolites for Each Pathway and Cross-correlations among the Pathways
Correlation between Enzymes and Metabolites for Each Pathway and Cross-correlations among the Pathways
Reference
Copyright © 2026 Creative Proteomics. All rights reserved.