Sinalbin (P-hydroxybenzyl Glucosinolate) Analysis Service
Submit Your InquiryWhat is Sinalbin?
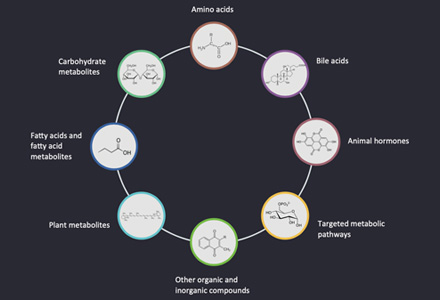
Sinalbin, also known as P-hydroxybenzyl glucosinolate, is a prominent member of the glucosinolate family of secondary metabolites. Glucosinolates are sulfur-containing compounds that are widely distributed in plants, particularly in the Brassicaceae family. Sinalbin is primarily found in various cruciferous vegetables, such as mustard, horseradish, radish, and white cabbage.
Structurally, sinalbin consists of a glucose molecule bound to a hydroxybenzyl group through a thioglucose linkage. This unique chemical structure imparts distinct properties to sinalbin, including its characteristic pungent aroma and taste.
Sinalbin serves several functions in plants, primarily as a defense mechanism against herbivores, pathogens, and environmental stresses. When the plant tissue is damaged, an enzyme called myrosinase hydrolyzes sinalbin, leading to the release of volatile compounds, such as isothiocyanates, nitriles, and thiocyanates. These compounds play a crucial role in deterring herbivores and defending the plant against microbial infections.
Apart from its role in plant defense, sinalbin has gained significant attention due to its potential health benefits. Research suggests that sinalbin and its breakdown products possess various biological activities, including antioxidant, antimicrobial, anti-inflammatory, and anticancer properties. These properties make sinalbin a subject of interest in the fields of food science, nutrition, and pharmaceutical research.
Analyzing sinalbin and understanding its distribution, biosynthesis, and metabolism provide valuable insights into the nutritional value of cruciferous vegetables, as well as their potential applications in functional foods, dietary supplements, and drug development.
Creative Proteomics offers a range of analytical services to support sinalbin analysis, enabling you to explore its properties, applications, and health-related implications.
-Analysis-Service-1.png) Structure of sinalbin (P-hydroxybenzyl glucosinolate)
Structure of sinalbin (P-hydroxybenzyl glucosinolate)
Sinalbin Analysis in Creative Proteomics
Sinalbin Quantitative Analysis
Accurate determination of sinalbin content is crucial for assessing its presence and concentration in various samples. Several analytical techniques are employed for quantitative sinalbin analysis, including:
- High-performance liquid chromatography (HPLC): HPLC coupled with UV or diode array detection (DAD) allows for sensitive and selective quantification of sinalbin in complex matrices.
- Gas chromatography (GC): GC combined with flame ionization detection (FID) or mass spectrometry (MS) offers precise analysis of sinalbin in volatile samples or after derivatization.
- Liquid chromatography-mass spectrometry (LC-MS): LC-MS techniques provide enhanced sensitivity and specificity for sinalbin analysis, enabling accurate quantification even at low concentrations.
Sinalbin Structural Identification and Confirmation
Understanding the biological activity and potential applications of sinalbin and its derivatives requires determining their structures. For structural identification, various techniques are used, including:
- Nuclear magnetic resonance spectroscopy (NMR): NMR spectroscopy, notably 1D and 2D NMR techniques, enables for the explanation of sinalbin's chemical structure, including the arrangement of atoms and functional groups.
- Mass spectrometry (MS): MS techniques such as electrospray ionization (ESI) or matrix-assisted laser desorption/ionization (MALDI) combined with high-resolution mass analyzers provide information on the molecular weight, fragmentation pattern, and existence of any modifications or adducts.
Sinalbin Metabolism Studies
Understanding sinalbin's bioavailability, transformation, and potential health implications requires research into its metabolism in living beings. Metabolism research entails extracting, purifying, and quantifying sinalbin and its metabolites from bsiological samples. Techniques typically used in sinalbin metabolism research include:
- Sample preparation techniques: These include solid-phase extraction (SPE), liquid-liquid extraction (LLE), and enzymatic hydrolysis to release sinalbin from its glucosinolate precursor.
- Chromatographic separation: Techniques such as HPLC or LC-MS are utilized for the separation and quantification of sinalbin and its metabolites in biological samples.
- Metabolite identification: Mass spectrometry-based techniques, such as high-resolution MS, tandem MS (MS/MS), and metabolomics approaches, aid in the identification and characterization of sinalbin metabolites.
Technical Advantages
- With standardized laboratory operation specifications and advanced instrument platform, the experiment period is short and the quality is reliable.
- It has multiple targeted instrument platforms such as Waters Quattro Premier XE triple quadrupole liquid-mass spectrometer (MRM), Agilent 7890A-5975C gas-mass spectrometer (SIM), etc.
- With strict quality control (internal standard and external standard), and manual proofreading standard products, the standard R2>0.99
- With a professional biological information team and large-scale hardware, it can provide customers with personalized biological information analysis services.
Technical Route of Targeted Metabolomics of Sinalbin (P-hydroxybenzyl Glucosinolate)
-Analysis-Service-2.png)
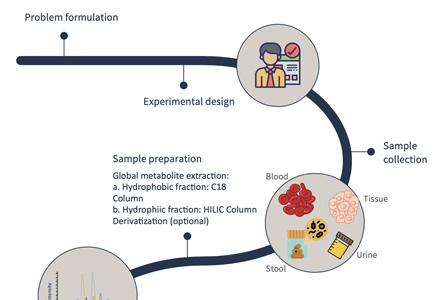
- Sample preparation
- Pre-test
- Make a standard curve
- Sample detection and result analysis
Extract metabolites, precipitate proteins, etc.
Optimize the corresponding parameters, establish the chromatographic and mass spectrometry analysis conditions of the tested substance, and create the MRM analysis method.
Configure a series of different concentration standards and make a standard curve.
According to the results of the sample detection, combined with the standard curve, the statistical analysis of the substance content in the sample is carried out.
Sample Preparation
| Sample type | Suggested sample quantity (per case) | Number of repetitions (recommended) |
|---|---|---|
| Plant tissue | ≥ 400 mg | Plant samples ≥ 3 cases / group Clinical samples ≥ 30 cases / group Animal samples ≥ 10 cases / group Cell samples ≥ 6 cases / group |
| Plant seed | ≥ 200 mg | |
| Blood (serum, plasma) | ≥ 200 μl | |
| Urine | ≥ 200 μl | |
| Animal tissue | ≥100 mg | |
| Cell | 107 | |
| Cell supernatant | ≥ 1ml | |
| Bacteria or fungi | Wet weight of sediment ≥ 200 mg | |
| Fermentation broth | ≥ 500 μl | |
| Other special samples | Please contact us for consultation | |
Feedback to Customers
Creative Proteomics will provide you with detailed technical reports, including
- Experimental steps
- Related mass spectrometry parameters
- Part of the mass spectrum picture
- Raw data
- Quantitative results of targeted metabolites
Project Cycle
- A standard experimental procedure takes about 1 ~ 4 weeks.