Taxadiene Analysis Service
Submit Your InquiryWhat are taxadiene?
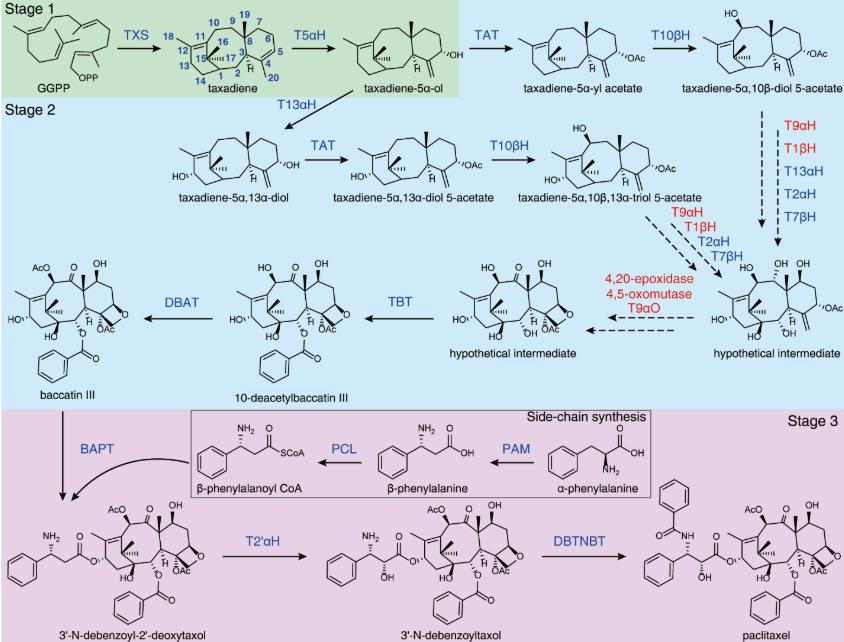
Taxadiene (C20H32), a key diterpene hydrocarbon, serves as the pivotal biosynthetic precursor for Taxol. Its biosynthesis begins with the cyclization of geranylgeranyl pyrophosphate (GGPP) catalyzed by taxadiene synthase (TXS), forming the unique tricyclic scaffold. As the first committed intermediate in Taxol biosynthesis, taxadiene quantification and structural analysis are critical for:
- Drug development: Monitoring flux through the Taxol biosynthetic pathway in Taxus species or engineered microbial systems.
- Metabolic engineering: Optimizing heterologous production of taxadiene in E. coli or yeast chassis.
- Quality control: Ensuring purity in commercial taxadiene batches for pharmaceutical intermediates.
Creative Proteomics delivers taxadiene analysis services tailored to accelerate research and industrial workflows.
Taxadiene Analysis Service by Creative Proteomics
Creative Proteomics provides a wide range of specialized services for Taxadiene analysis to ensure high-precision data back your research and development processes. Our service offerings include:
1. Taxadiene ldentification and Quantification
- Targeted GC-MS or LC-MS/Ms analysis for accurate quantification of taxadiene
- Absolute and relative quantification with internal standards
- Detection sensitivity at trace levels in plant extracts, microbial cultures, or engineered systems
2.Pathway Profiling and Metabolite Analysis
- comprehensive metabolic profiling to identify taxadiene biosynthetic intermediates
- Monitoring of taxadiene pathway fluxes using stable isotope labeling (SIL)
- Comparative analysis of engineered strains for metabolic optimization
3. Structural Characterization
- High-resolution MS and MS/MS analysis for taxadiene structural elucidation
- Fragmentation pattern analysis to determine molecular structure and confirm identity
4. Isotope Tracing Studies
- Quantifying metabolic rates by labelling paclitaxel and other paclitaxel metabolites reveals dynamic metabolic processes.
- Combined with mass spectrometry to track the binding of isotopically labelled compounds in metabolic pathways to determine metabolic rates and pathway activity.
5. Custom Assay Development
- Development and validation of customized taxadiene quantification assays
- Optimization of extraction and sample preparation protocols for diverse sample matrices
6. Data Interpretation and Report Generation
- Detailed, scientifically rigorous reports, including all relevant data, trends and analyses
- Assist in decision-making and further research
List of Taxadiene-Related Metabolites (including but not limited to)
| Detected Substance | Type | Metabolic Pathway |
|---|---|---|
| Taxadiene | Diterpene | Biosynthesis of paclitaxel from taxadiene in Taxus species |
| Taxol (Paclitaxel) | Terpenoid | Conversion from taxadiene via several enzymatic steps |
| 10-deacetylbaccatin III | Precursor to Paclitaxel | Intermediate in the paclitaxel biosynthetic pathway |
| Baccatin III | Diterpene | Intermediate leading to paclitaxel formation |
| 7-epi-taxol | Taxoid derivative | Side-product in the biosynthesis pathway |
| CYP450 Enzymes | Enzyme family | Key enzymes involved in hydroxylation steps in taxadiene metabolism |
| geranylgeranyl pyrophosphate(GGPP) | Precursor | Terpenoid synthesis precursors |
Techniques and Instrumentation for Taxadiene Analysis
 Bruker 400 MHz NMR spectrometer (Figure from Bruker)
Bruker 400 MHz NMR spectrometer (Figure from Bruker)
 7890B Gas Chromatograph (Figure from Agilent)
7890B Gas Chromatograph (Figure from Agilent)
 ISQ™ EC Single Quadrupole MS (Figure from Thermo Scientific)
ISQ™ EC Single Quadrupole MS (Figure from Thermo Scientific)
 Mass-Based 1290 Infinity II Preparative LC/MSD System (Figure from Agilent)
Mass-Based 1290 Infinity II Preparative LC/MSD System (Figure from Agilent)
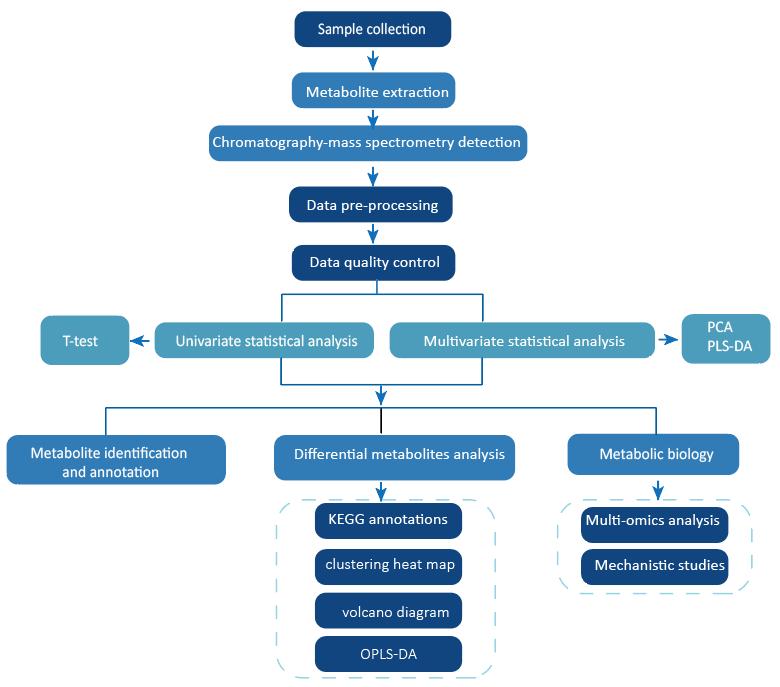
Workflow for Taxadiene Analysis Service
Why Choose Us?
Applications of Taxadiene Analysis
|
|
Biotechnology and Synthetic Biology Investigating microbial or plant-based systems engineered for the biosynthesis of taxadiene and its derivatives. |
|
Agricultural Science Understanding the genetic and environmental factors that influence taxadiene production in Taxus species or other plants |
|
|
Pharmacology and Drug Development Studying the pathways that lead to paclitaxel production and optimizing taxadiene biosynthesis for pharmaceutical applications. |
|
Academic Research Map taxadiene's role in plant defense mechanisms |
Sample Requirements for Taxadiene Analysis Assay
| Sample Types | Minimum Quantity | Required Conditions |
|---|---|---|
| Plant Tissues | 500 mg | Freeze-dried or fresh samples, stored at -80°C |
| Microbial cultures | 10-50 mL | Liquid cultures grown under controlled conditions |
| Plant extracts | 2-5 mL | Extracts should be in organic solvents like methanol or ethanol |
| Cell cultures (suspension) | 1-5 mL | Harvest cells in mid-log phase, flash-frozen |
- Q: How long does a standard taxadiene quantification take?
- A: 24–48 hours for GC-MS quantification; 72 hours for full pathway flux analysis.
- Q: Can I submit lyophilized samples?
- A: Yes, but reconstitute in 500 µL hexane before shipping at -20℃.
- Q: What's the LOD for taxadiene in yeast cultures?
- A: 0.5 ng/mL (matrix-matched calibration applied).
- Q: Do you recommend technical replicates?
- A: Yes—we include triplicate runs at no extra cost.
- Q: Can I receive raw data files?
- A: Yes, .RAW (Thermo), .D (Agilent), and .Wiff (Sciex) formats provided.
- Q: Do you analyze taxadiene epoxides?
- A: Yes, via LC-QTOF (requires ≥50 µL sample).
- Q: How is data confidentiality handled?
- A: All data is encrypted, and projects are assigned anonymized IDs.
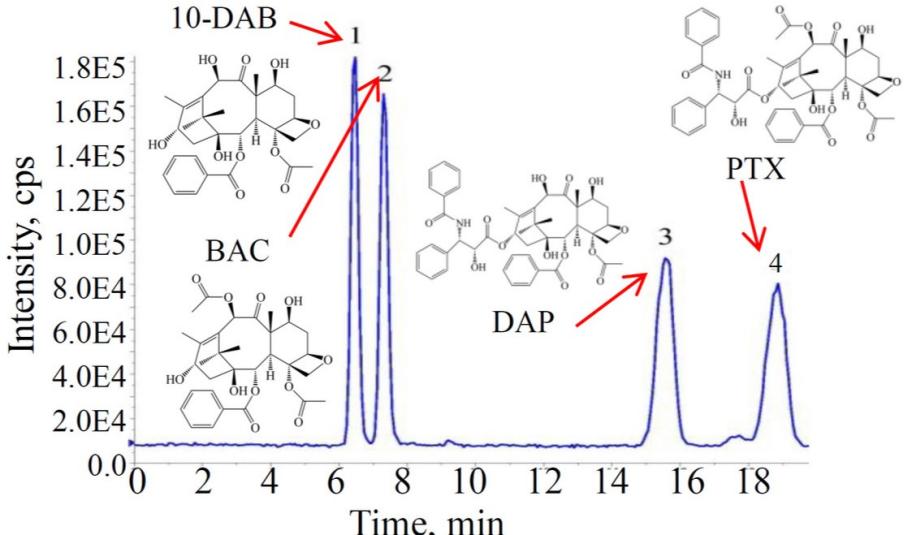 Chromatogram displaying intensity vs. time for 10-DAB, BAC, DAP, and PTX compounds with cps values ranging from 1.8E5 to 2.0E4 (Figure from Chunna Yu et al., 2020)
Chromatogram displaying intensity vs. time for 10-DAB, BAC, DAP, and PTX compounds with cps values ranging from 1.8E5 to 2.0E4 (Figure from Chunna Yu et al., 2020)
Case. Characterization and heterologous reconstitution of Taxus biosynthetic enzymes leading to baccatin III
Background:
- Gene expression analysis of candidate genes and paclitaxel biosynthetic genes.
- Metabolite extraction and LC-MS analysis.
- This analysis aims to identify two key deletion genes in the paclitaxel synthesis pathway, TOT and T9αH, to reveal a novel mechanism by which plant cells catalyze the formation of oxetane structures.
Samples:
- Different tissues of Picea abies (needles, bark and roots of females (F) and males (M)), Nicotiana benthamiana, freeze-dried tissues.
- Three biological replicates were included for each line, with six plants for each biological replicate
Technical methods procedure:
- Expression levels of candidate genes and paclitaxel biosynthesis genes were analyzed based on RNA-seq data.
- Taxus mairei, Nicotiana benthamiana and insect cells were freeze-dried.
- Metabolites were extracted by sonication of freeze-dried tissues with 1 mL MeOH for 20 min at room temperature.
- Centrifugation to obtain supernatant
- Liquid chromatography-electrospray ionization-tandem mass spectrometry (LC-ESI-MS/MS) was used for qualitative analysis of metabolites.
- Metabolites were quantified using standard curves.
- Concentration determination of paclitaxel metabolite content using an ion chromatography system.
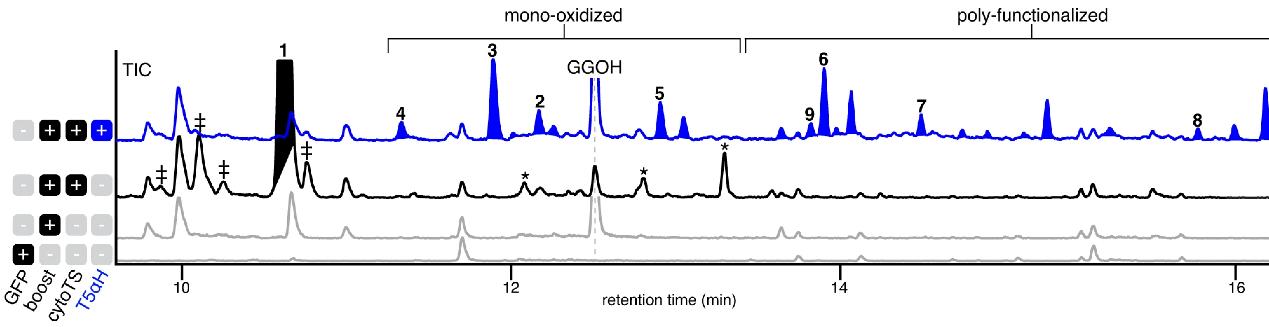
Results
- Screening and validation of oxidase synthesis genes in taxadiene metabolism.
- Determination of changes in the content of intermediates in the paclitaxel synthesis pathway.
- Identification of a bifunctional cytochrome P450 enzyme [paclitaxel oxetane enzyme 1 (TOT1)] in Taxus mairei that catalyzes an oxidative rearrangement in paclitaxel oxetane formation.
- The biosynthetic pathway of baccatin III was successfully artificially reconstructed in tobacco by co-expressing other known genes involved in baccatin III synthesis.
References
- Bin Jiang et al., Characterization and heterologous reconstitution of Taxus biosynthetic enzymes leading to baccatin III.Science383,622-629(2024). https://doi.org/10.1126/science.adj3484
- Li, J., Mutanda, I., Wang, K. et al. Chloroplastic metabolic engineering coupled with isoprenoid pool enhancement for committed taxanes biosynthesis in Nicotiana benthamiana. Nat Commun 10, 4850 (2019). https://doi.org/10.1038/s41467-019-12879-y.
- Jin-Yi Wang, Zheng-Yu Huang. et al. Facile biosynthesis of taxadiene by a newly constructed Escherichia coli strain fusing enzymes taxadiene synthase and geranylgeranyl pyrophosphate synthase.Process Biochemistry 122, 2 (2022). https://doi.org/10.1016/j.procbio.2022.09.005